10 pts total
Problem 1
A - 1pt
load this dataset on scapula morphology in Pan, Homo and Gorilla, and the Dikika fossil child. Details on this data can be found in Alemseged et al, 2006
scap <- read.table("https://stats.are-awesome.com/datasets/DIK-scap.txt", header=TRUE)B - 1pt
Calculate the natural log of all the numeric variables
# log all the measurements, excluding the first column
scap[,-1] <- log(scap[,-1])C - 1pt
Make a subset of the data including only the extant species (G, H, or P, representing Gorilla, Homo and Pan), excluding the Dikika fossil (D).
subset_scap <- dplyr::filter(scap, Taxon != "D")D - 2pt
Perform a discriminant function analysis on the data. Which variable has the strongest loading on LD1?
library(MASS)##
## Attaching package: 'MASS'## The following object is masked from 'package:dplyr':
##
## selectDFA <- lda(x=subset_scap[,-1], grouping=subset_scap$Taxon)
DFA## Call:
## lda(subset_scap[, -1], grouping = subset_scap$Taxon)
##
## Prior probabilities of groups:
## G H P
## 0.3417722 0.2911392 0.3670886
##
## Group means:
## SSFB TOTB SSFL ISFB MOL SPL ISFL PSSFB
## G 4.213874 4.816231 4.201309 4.225263 4.522603 4.713122 4.621712 3.854627
## H 3.260568 4.345349 3.657431 4.106738 3.980998 4.178805 4.049559 3.072758
## P 3.897599 4.518186 3.918844 3.867818 4.182304 4.398842 4.423187 3.408756
## PISFB GFL GFB
## G 4.210353 3.391814 3.005870
## H 4.100065 3.030641 2.631709
## P 3.635822 3.088464 2.741510
##
## Coefficients of linear discriminants:
## LD1 LD2
## SSFB -10.3161467 -4.2460350
## TOTB -2.8354310 -0.7961162
## SSFL 11.1350006 10.0673675
## ISFB -3.2755157 12.0736770
## MOL -4.4411419 -17.8401790
## SPL 0.9830035 6.3123756
## ISFL -4.9539689 5.1112446
## PSSFB 0.6174285 -4.2105436
## PISFB 13.1771010 -11.5279013
## GFL -1.2594441 -0.4065495
## GFB 1.7064927 3.4084295
##
## Proportion of trace:
## LD1 LD2
## 0.8095 0.1905E - 1 points
PISFB has the strongest loading
F - 3 pt
Make a beautiful plot (using ggplot) of LD1 versus LD2. Hint: to get the LD scores, use the predict() function on your discriminant function analysis model object, as shown in class.
library(ggplot2)
forPlot <- data.frame(predict(DFA)$x, taxon=subset_scap$Taxon)
LDAscatter <- ggplot(forPlot, aes(x=LD1, y=LD2)) + geom_point(aes(color=taxon))
LDAscatter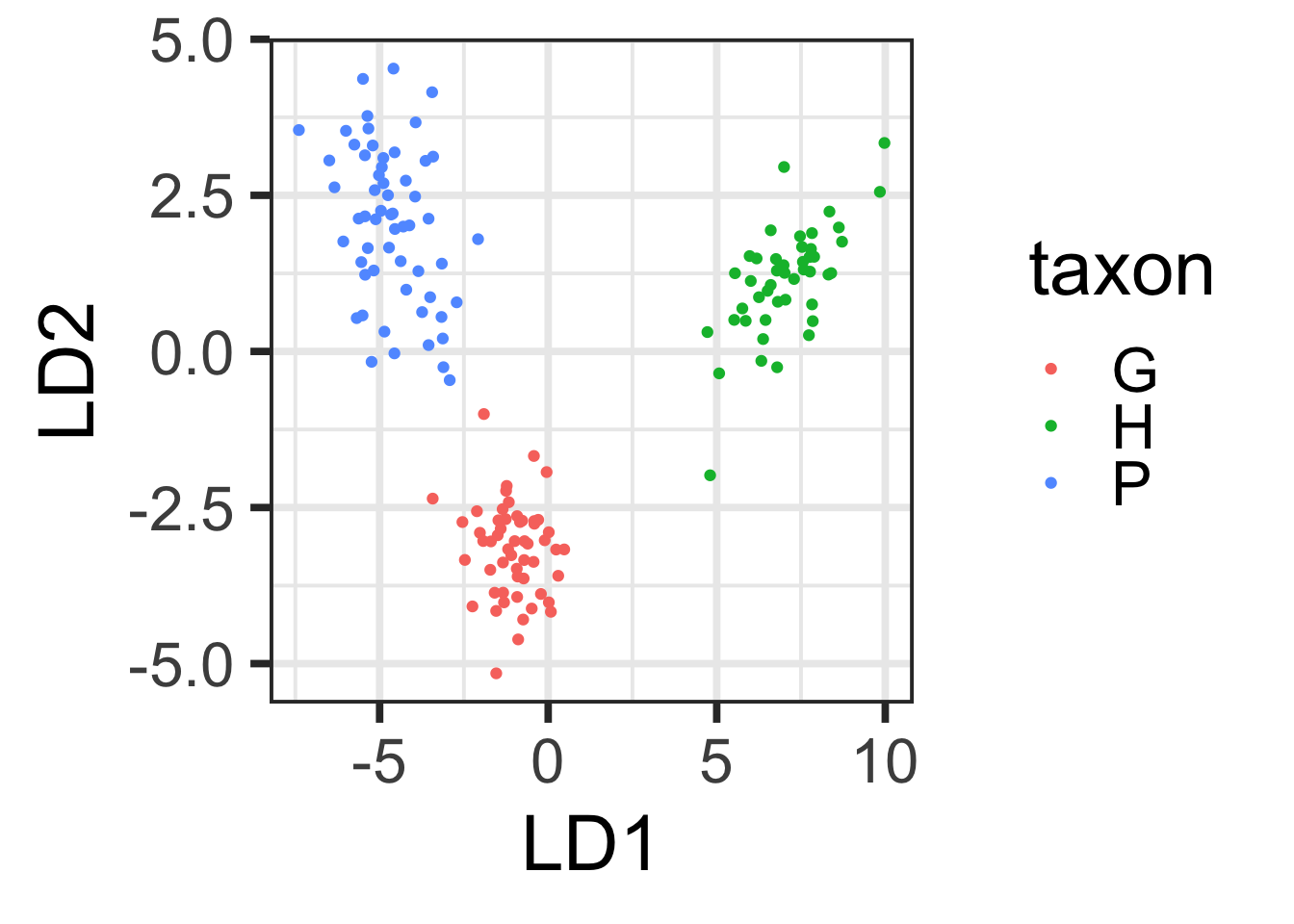
F - 1 bonus pt
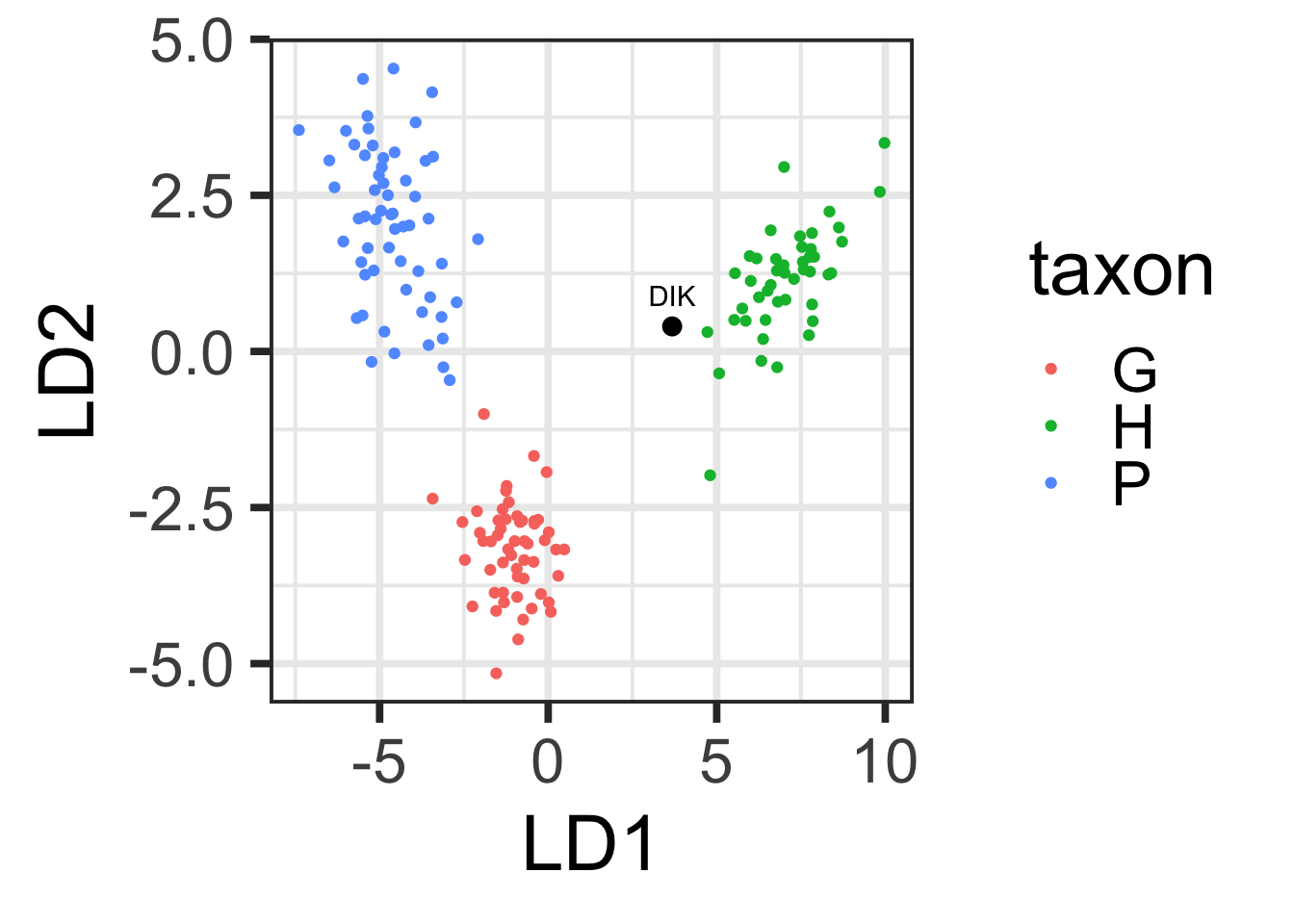
Plot the Dikika fossil on the DFA plot.
dikika <- dplyr::filter(scap, Taxon=="D")
DIK_dfa <- predict(DFA, newdata = dikika[,-1])
DIK_plot <- data.frame(DIK_dfa$x, label="DIK")
LDAscatter +
geom_point(data=DIK_plot,size=3) +
geom_text(data=DIK_plot,aes(x=LD1, y=LD2 + 0.5, label=label))