15 total points for homework
Question 1
- 2pts shapiro test
- 2pts histogram
- 1pt mean and median
- 1pt why mean and median differ
Part A
Note that columns 1 thorough 4 are the numeric columns in the iris dataset.
for(i in 1:4){
print(shapiro.test(iris[,i])$p.value)
}## [1] 0.01018116
## [1] 0.1011543
## [1] 7.412263e-10
## [1] 1.680465e-08Part B
library(ggplot2)
qplot(iris$Sepal.Length) ## Warning: `qplot()` was deprecated in ggplot2 3.4.0.
## This warning is displayed once every 8 hours.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.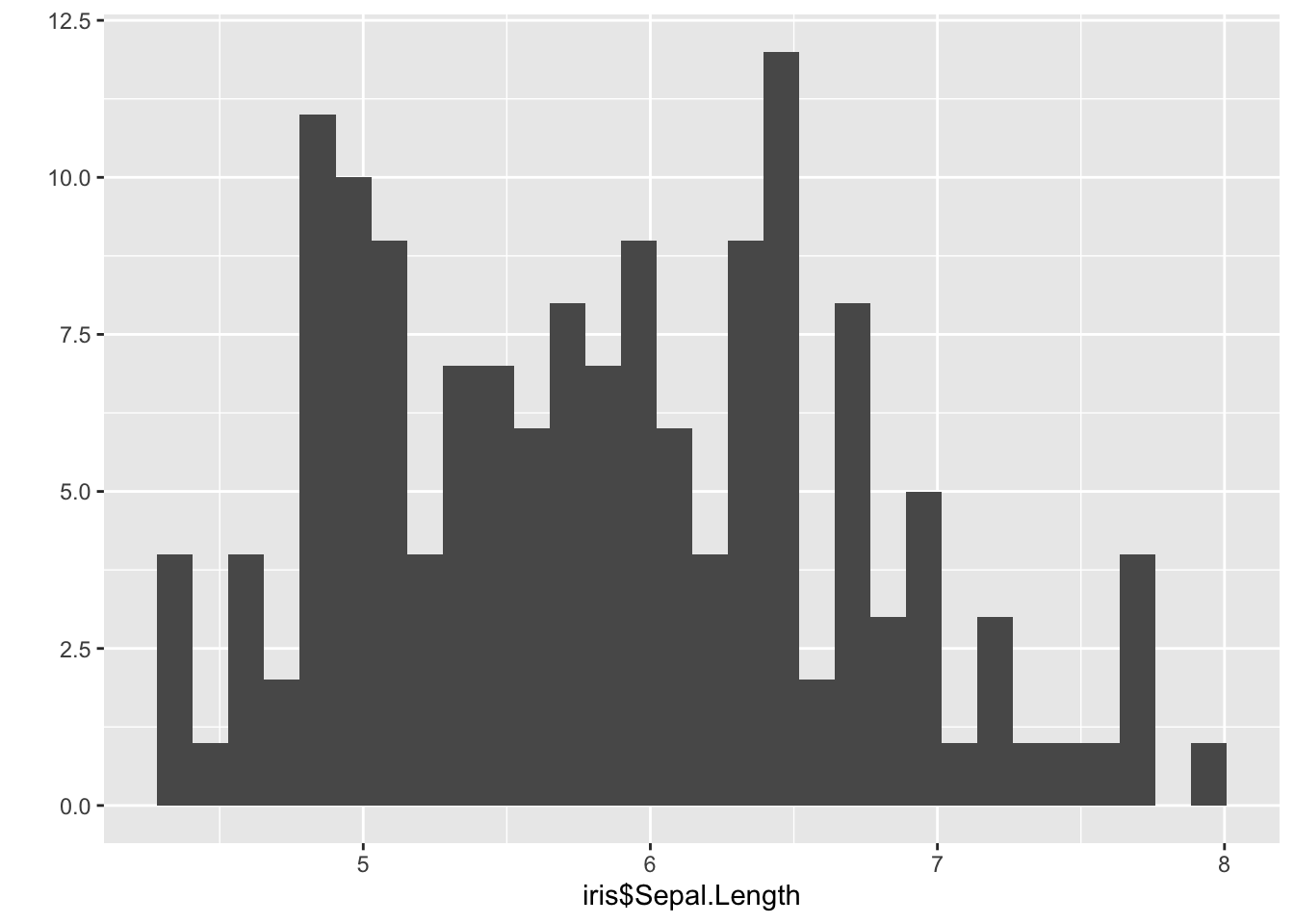
qplot(iris$Sepal.Width) ## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.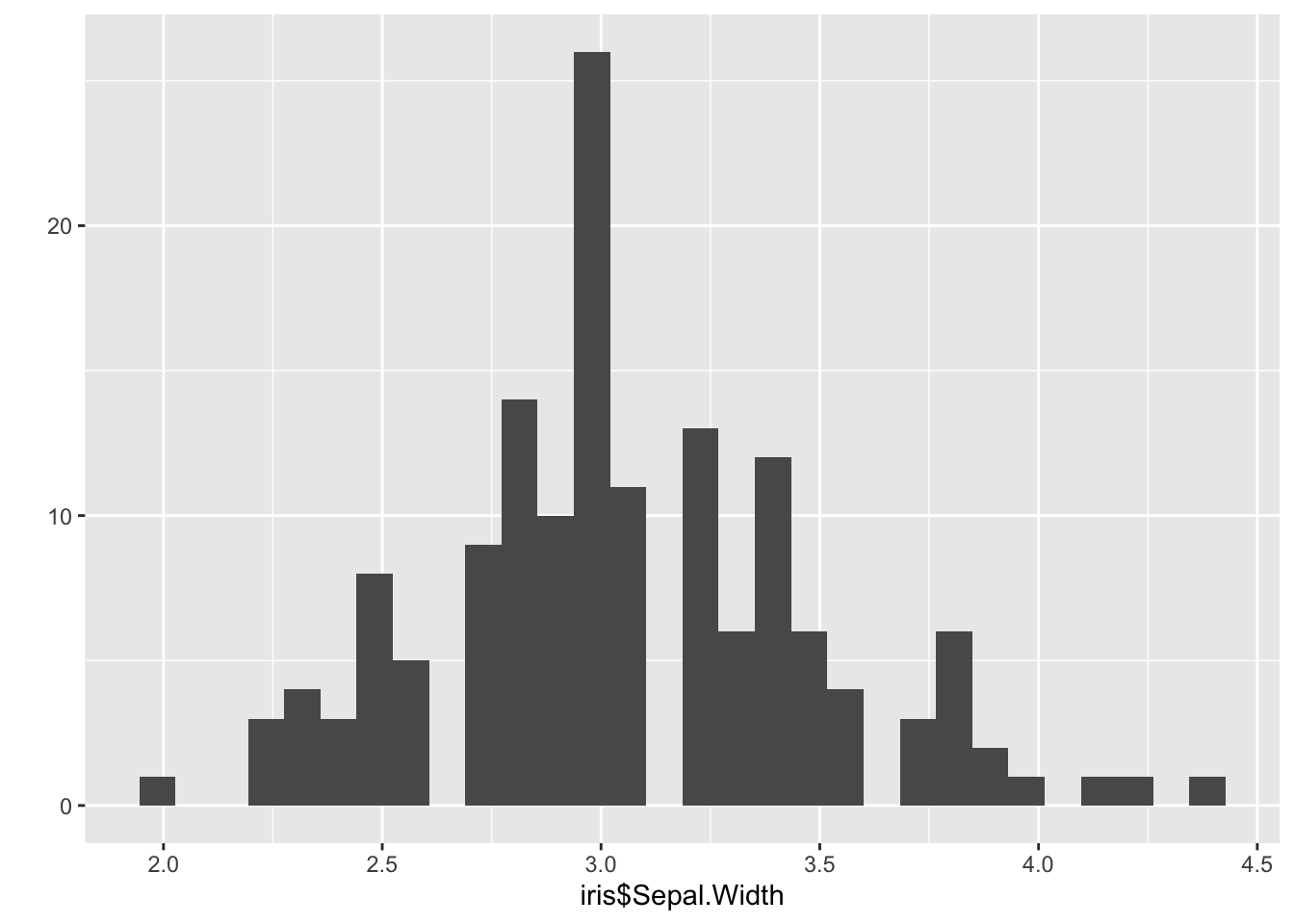
qplot(iris$Petal.Length) ## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.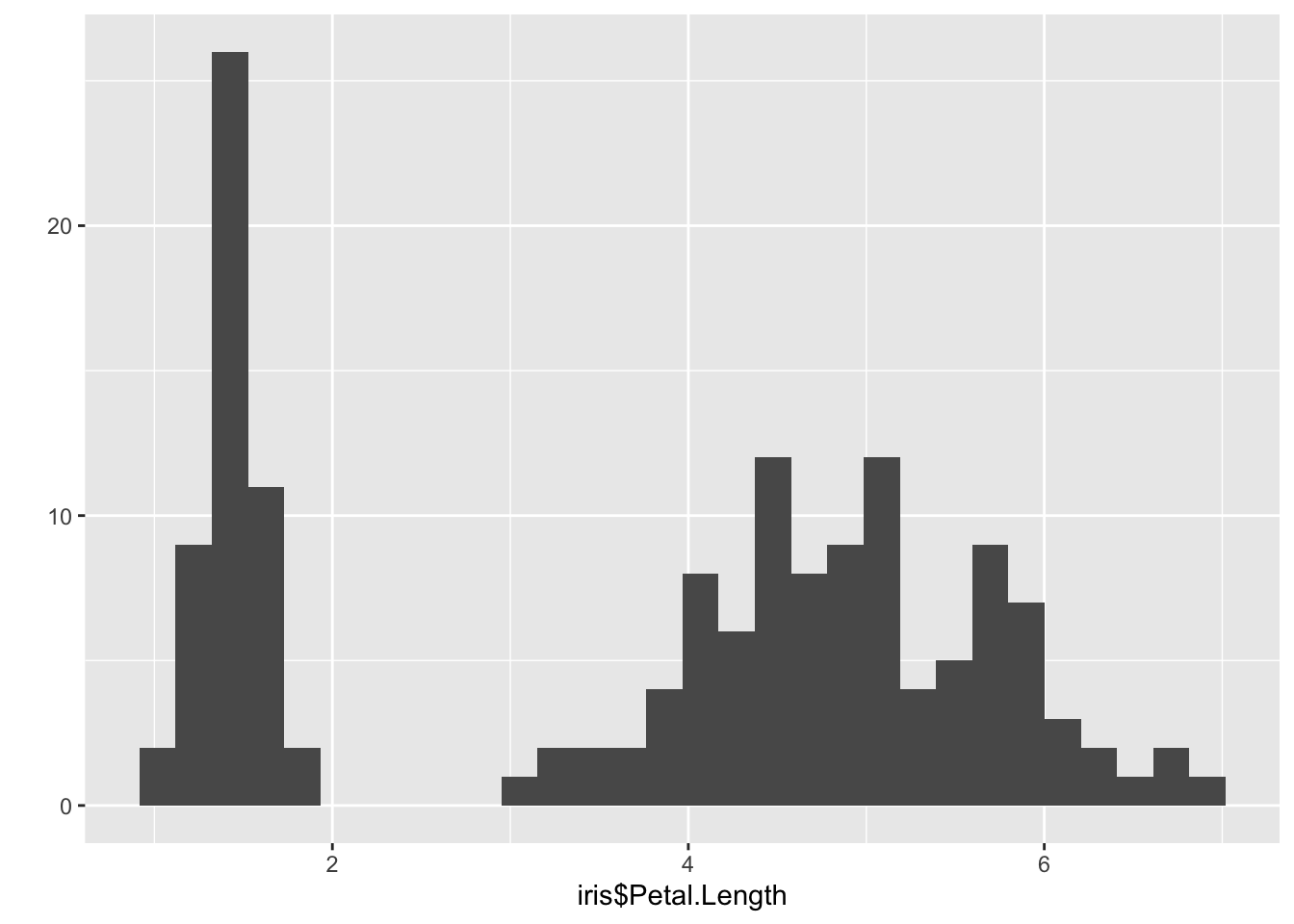
qplot(iris$Petal.Width) ## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.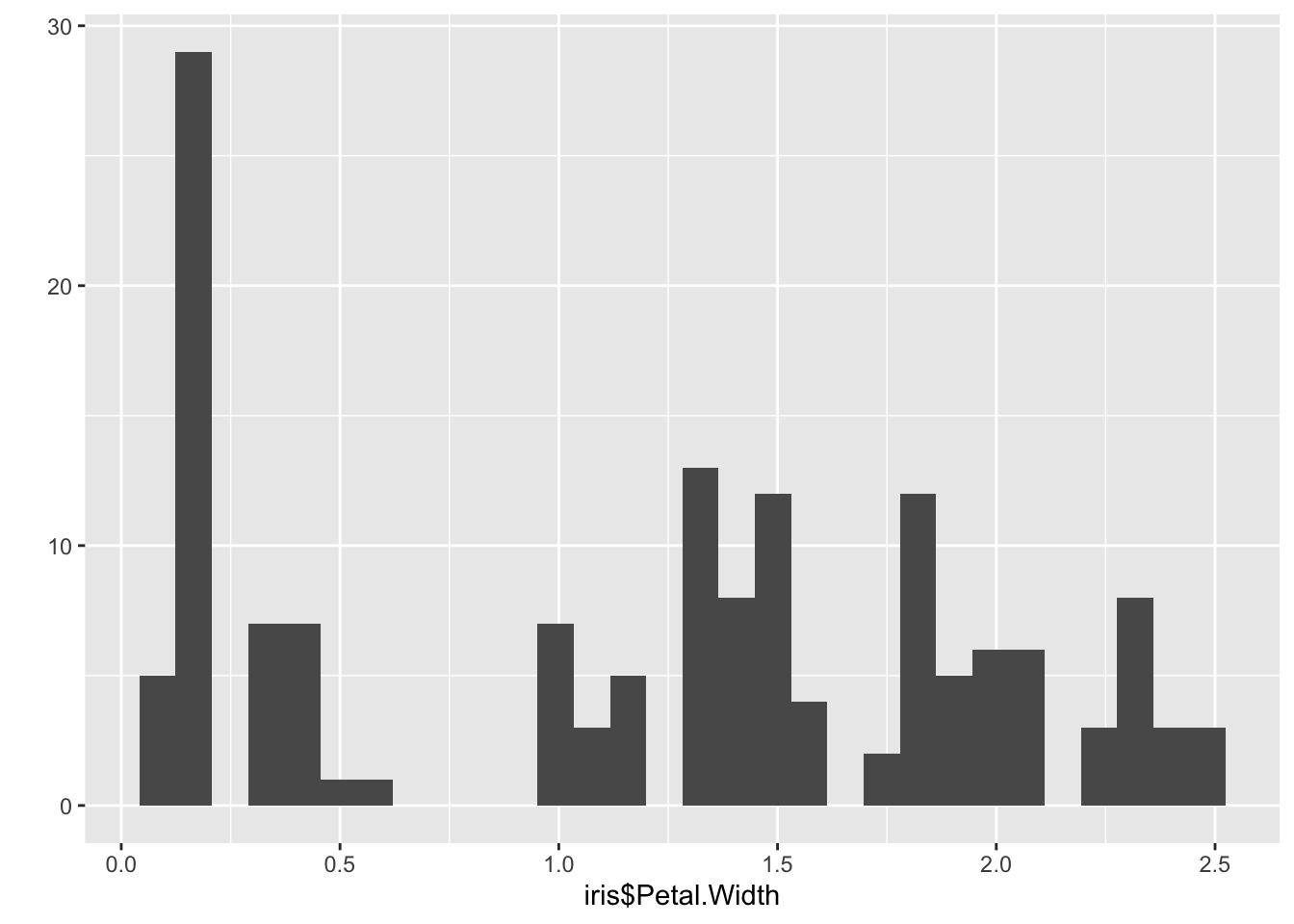
Part D
mean(iris$Sepal.Length^2)## [1] 34.82567median(iris$Sepal.Length^2)## [1] 33.64Part E
These values are different because the distribution of iris$Sepal.Length^2 is asymmetric, as shown in the below histogram.
qplot(iris$Sepal.Length^2)## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.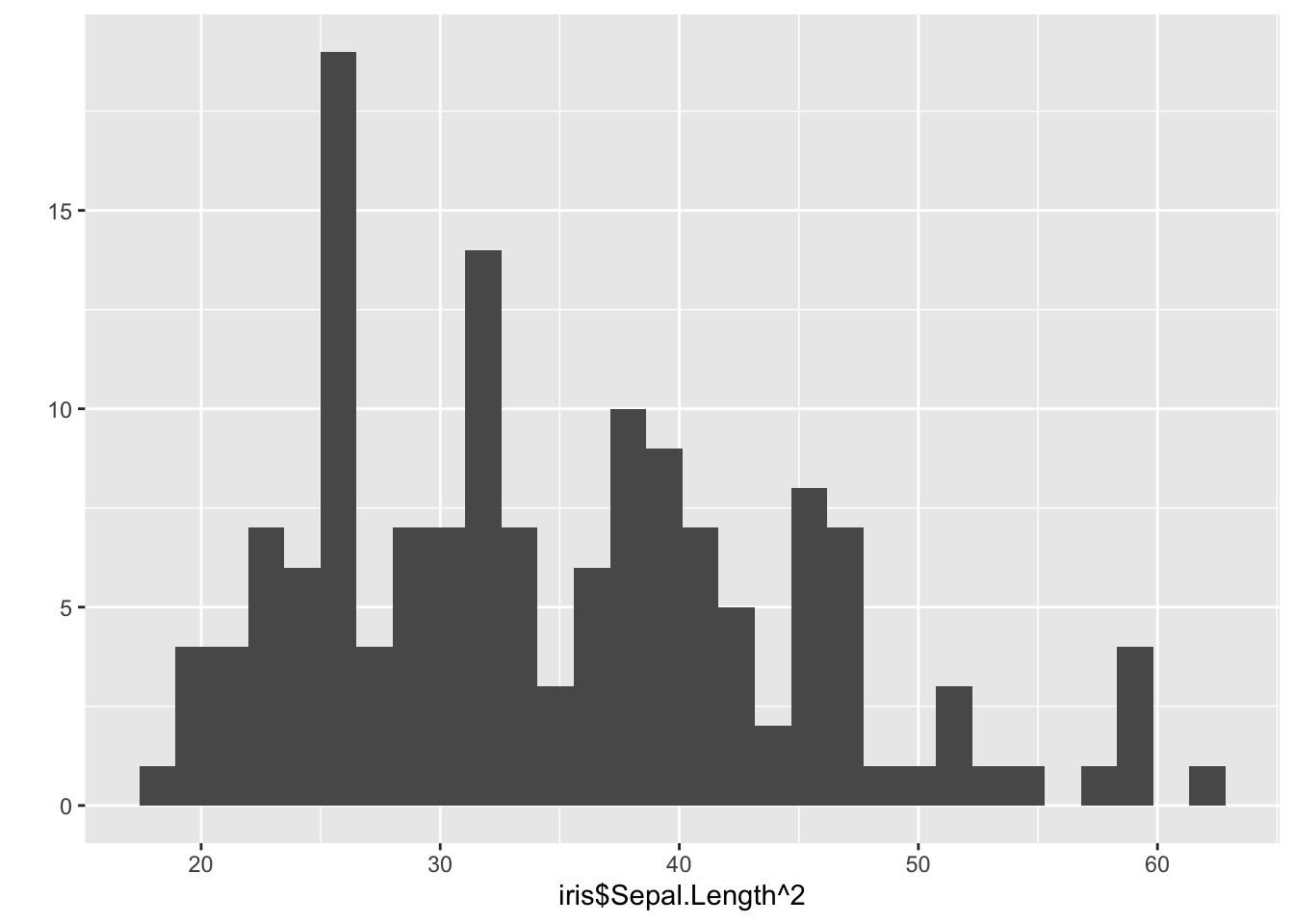
The distribution has a long right tail, which heavily influences the mean, pulling it towards the larger values. The median is less affected by extreme values.
Question 2
- 4 pts for the for loop / lapply correcly reading in all files
- 1 pt for correctly reporting average
- 2 pts histogram
- 2 pts answer question is it normal
Part A
filez <- list.files("/Path/to/folder", full=TRUE)Set up and run a for() loop
# create empty vector, to be filled up with the averages
proportions_of_G <- rep(NA,100)
#loop through index numbers of files
for(i in 1:length(filez)){
#read file in
df <- read.table(filez[i], header=T, sep=",")
#calculate the proportion of G
propG <- sum(df$base == "G") / length(df$base)
#update the results vector with the proportion from current file
proportions_of_G[i] <- propG
}Alternatively, you could do it with sapply() and a custom function. Note: the difference between lapply() and sapply() whether you get a list or a vector as a result (l stand for list).
doIt <- function(filename){
df <- read.table(filename, header=T, sep=",")
propG <- sum(df$base == "G") / length(df$base)
return(propG)
}
proportions_of_G <- sapply(filez, doIt)After running the for loop, we can report the average proportion of G across all files
mean(proportions_of_G)## [1] 0.2491692Part B
library(ggplot2)
ggplot(mapping=aes(x=proportions_of_G)) +
geom_histogram(bins=10) +
labs(x="Proportion of G",
title="Proportions of Guanine across 100 genetic sequences")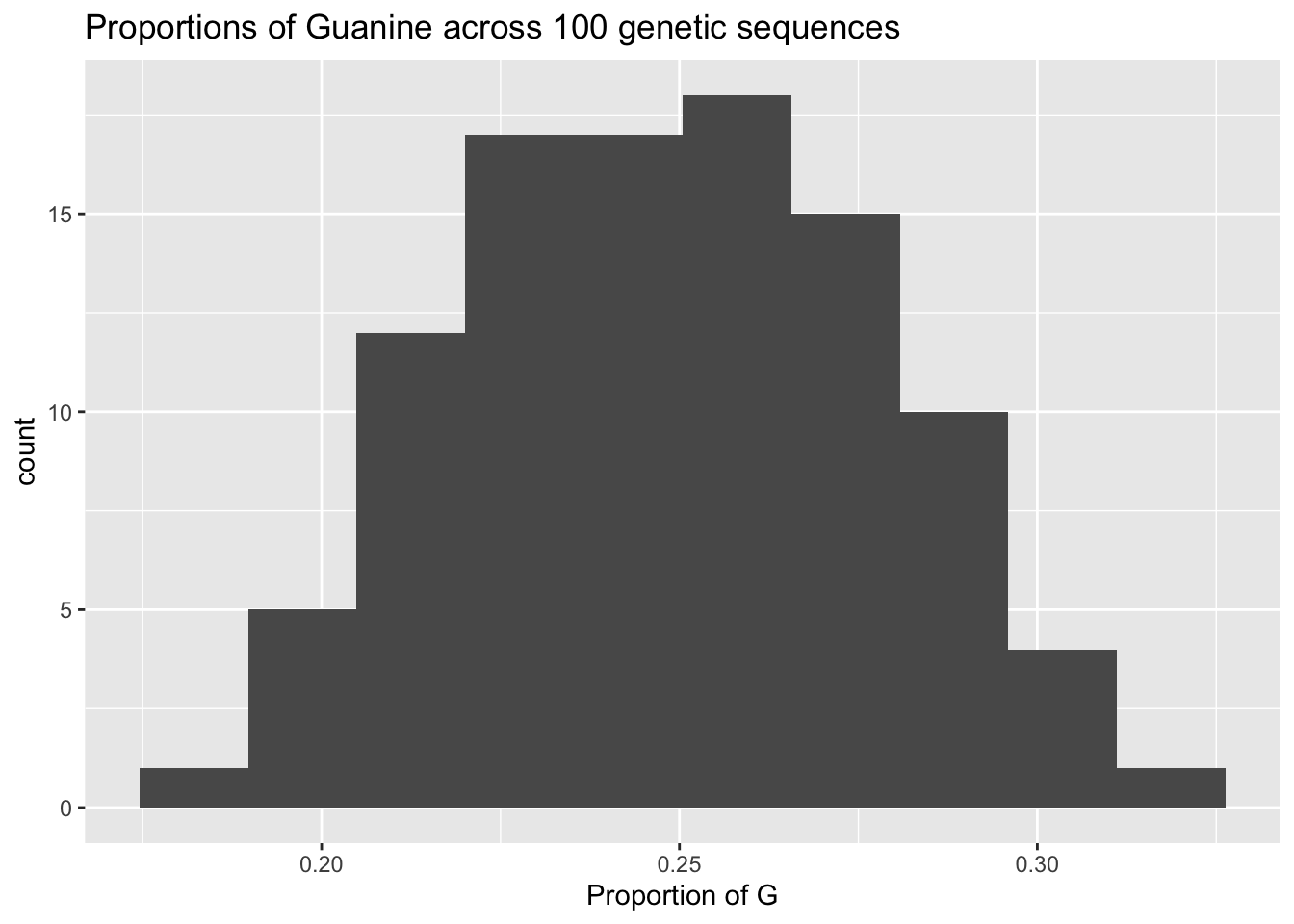
Part C
This is a decently normal distribution. It is fairly symmetrical around the mean…its pretty unimodal. If we want to put a number on our intuition, the shapiro.test() agrees it is normal.
shapiro.test(proportions_of_G)##
## Shapiro-Wilk normality test
##
## data: proportions_of_G
## W = 0.99236, p-value = 0.8469